 The PolyPharma app is currently getting some major new content added to it, namely a bunch of new models for toxicity. The models are derived from measurements made on the EPA’s Tox21 collection, recently published in nature, and released via PubChem.
The PolyPharma app is currently getting some major new content added to it, namely a bunch of new models for toxicity. The models are derived from measurements made on the EPA’s Tox21 collection, recently published in nature, and released via PubChem.
This nice little development lent itself immediately to the treatment that was recently applied to the ChEMBL dataset, which involved chopping up all of the different target groups, and selecting categories for merging into model-ready source collections. The total number of targets and off-targets was more than 1800, of which a selection of them were picked for inclusion in the PolyPharma app. The EPA Tox21 measurements on the other hand make up a much smaller number of targets – 29 to be exact – and these can be pulled down out of PubChem with a relatively simple script. The measurements have already been classed as either active (i.e. bad) or inactive, which makes life easier – no need to figure out a threshold prior to sticking into a Bayesian model.
Generally speaking, the structures that went into the dataset were of high quality, though a number of corrections are still necessary: washing out salts and adducts, a small number of egregious impossible molecules, and quite a lot of inorganics/organometallics with an inconsistent approach to dealing with non-boring bond types, some of which cannot be represented using normal Molfiles anyway. These had to be fixed manually, which was quite labour intensive.
Long story short, using ECFP6 fingerprints to generate Bayesian models resulted in quite agreeable statistics:
| Target | ROC | Actives/Size |
| ATAD5 | 0.8955282143671504 | 372/9819 |
| DT40 Rev3 | 0.8358695790539805 | 2470/8564 |
| DT40 WT | 0.8481449024919616 | 2489/8964 |
| P53-bla | 0.8671812374075394 | 659/9403 |
| RE-bla | 0.8199006118936651 | 1131/7322 |
| HSE-bla | 0.7807963830271731 | 497/8376 |
| romatase | 0.8564348950930594 | 378/7846 |
| mitochondria toxicity | 0.8975806303770444 | 1229/7958 |
| hR-luc | 0.897262258206716 | 1058/8878 |
| R-bla agonist | 0.8614046773463251 | 332/9299 |
| R-bla antagonist | 0.8637624381527989 | 647/8316 |
| R-MDA-luc agonist | 0.7773459658086779 | 414/10133 |
| R-MDA-luc antagonist | 0.8322296191717147 | 468/8269 |
| ER-BG1-luc agonist | 0.7402058726456726 | 1017/8340 |
| ER-BG1-luc antagonist | 0.868486076379126 | 472/8666 |
| ER-bla agonist | 0.8224802258757133 | 560/9545 |
| ER-bla antagonist | 0.8269233837880261 | 432/8297 |
| FXR-bla agonist | 0.7711240470264435 | 118/8674 |
| FXR-bla antagonist | 0.8486710970456315 | 255/7833 |
| GR-bla agonist | 0.8291952546091927 | 211/9256 |
| GR-bla antagonist | 0.8502523415129595 | 452/8094 |
| PPAR-delta-bla agonist | 0.8070517362777032 | 112/8274 |
| PPAR-delta-bla antagonist | 0.7698014607724666 | 92/7937 |
| PPAR-gamma-bla agonist | 0.8513509469037785 | 277/8898 |
| PPAR-gamma-bla antagonist | 0.8354619508727978 | 428/7610 |
| TR-beta-luc agonist | 0.7290186160244048 | 64/9422 |
| TR-beta-luc antagonist | 0.826791127196751 | 419/7391 |
| VDR-bla agonist | 0.6806460193156809 | 22/8437 |
| VDR-bla antagonist | 0.811523182164013 | 83/7699 |
So far so good. The next step is to actually do something with these models, in the same way as we have been for the ChEMBL extracts. Currently this consists of two pathways: one is creating detailed reports that can be perused and scrolled through when applied to individual molecules, such as discontinued drugs, and the other is to add the functionality to PolyPharma, so that anyone with an iThing can try it out.
All of the targets have been added to the mix, and there is now a default Profile called “Toxicity” which focuses entirely on these newly added models:
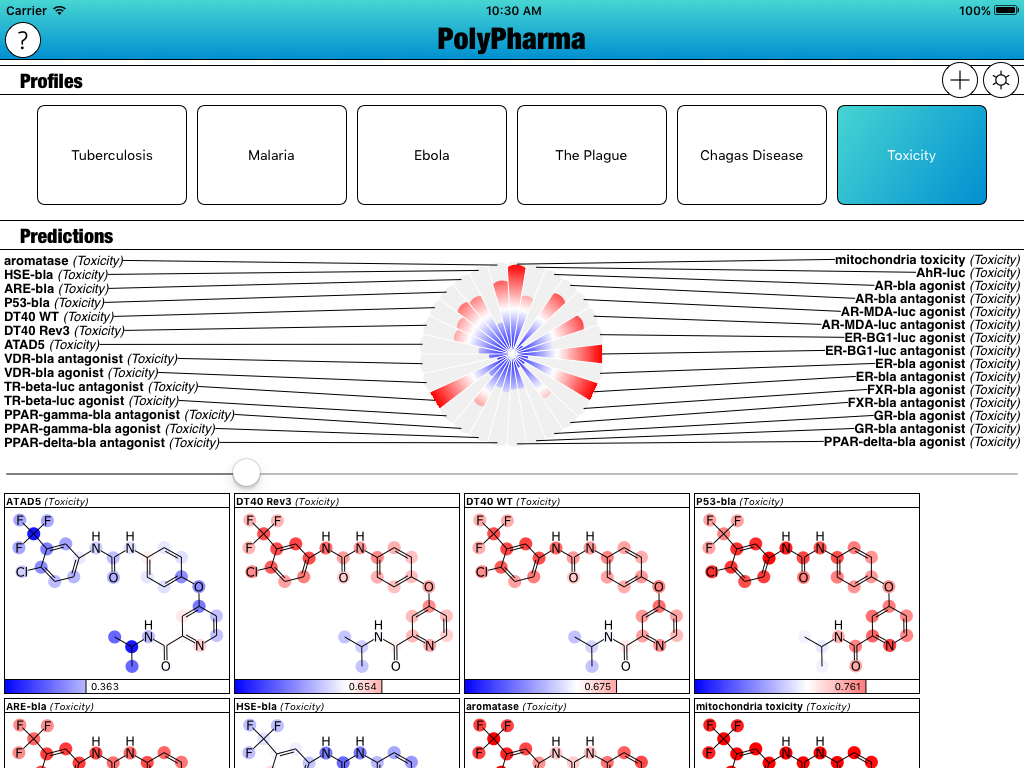
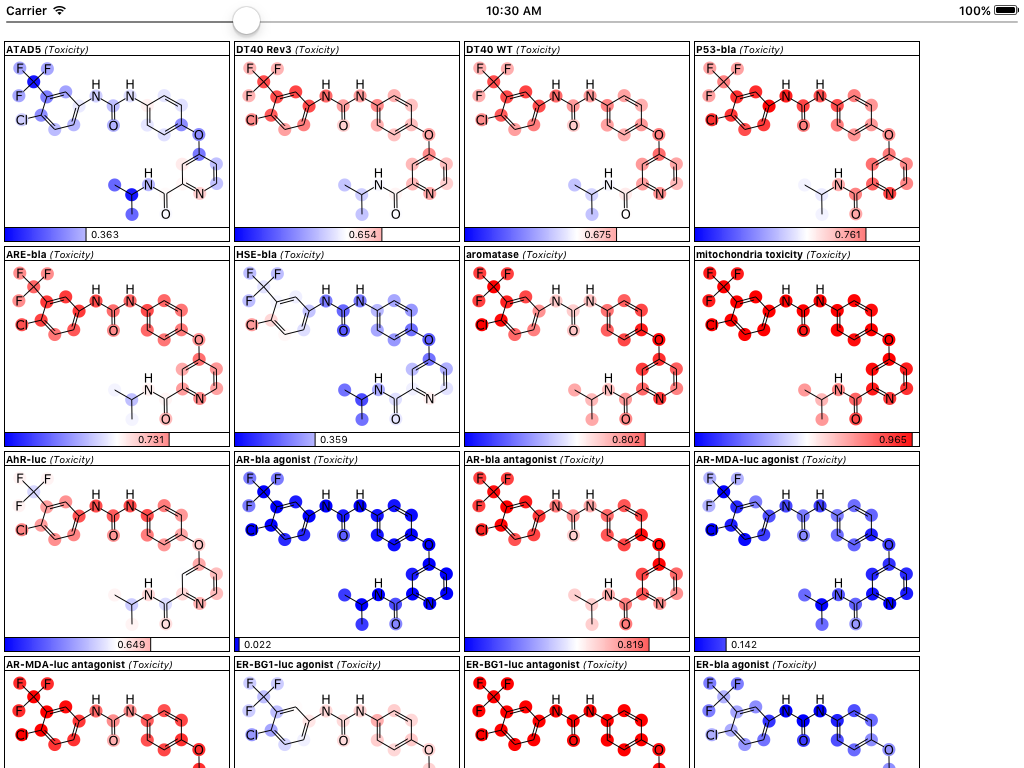
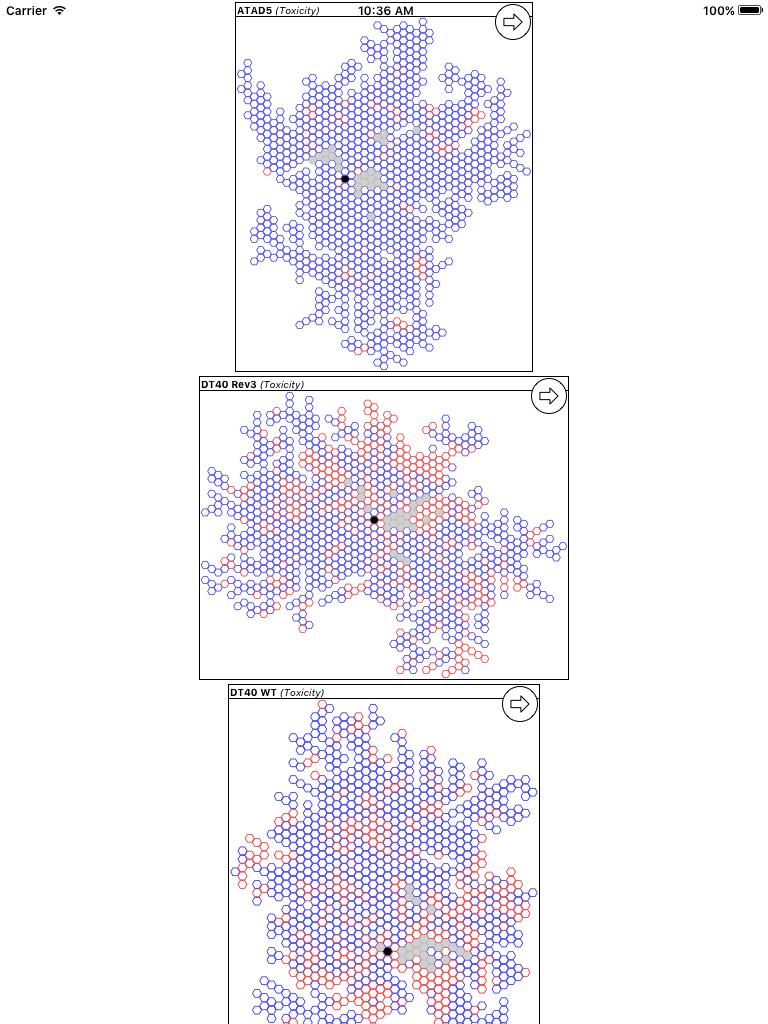
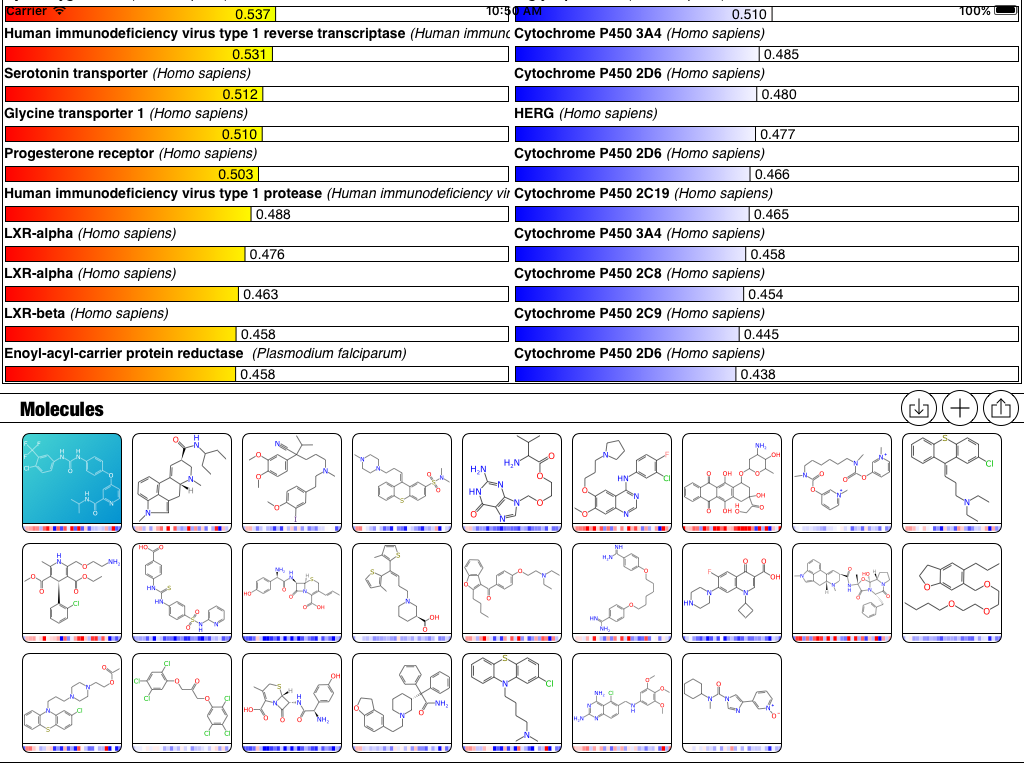
The updated version of the app hasn’t been submitted to the AppStore just yet, but it will be shortly, after a bit more testing. Stay tuned for the next version!