Large Language Models (LLMs or, in the parlance of our times, “AI”) have some potential value for reversing the translation of chemical reaction experiments into scientific English, into something more digitally friendly. Or put another way, there is an enormous amount of chemical reaction data that exists only as text, and if there was a less labour-intensive way to extract it, we would be much better off.
This is a series of articles about reaction prediction. The summary overview and table of contents can be found here. The website that provides this functionality is currently in a closed beta, but if you are interested in trying it out, just send an email to alex.clark@hey.com and introduce yourself.
In this article we’ll take a look at a prototype that is in a much earlier stage than the tools described in the previous sections. At the time of writing (June 2025) it is very much a skunkworks project.
First of all my thoughts on the AI hype-obsession that has engulfed the globe are not very flattering: most of the use cases that pop up in every day conversations could be charitably described as exorbitant parlour tricks. It’s a bit like having a kitten that talks – the novelty is well and truly off the charts, but eventually everyone has to face up to the reality that the talking cat doesn’t really have anything particularly interesting to say. Nonetheless, anyone with a grasp of the core technology can imagine that there are use cases that are very much appropriate. A model that is able to simulate language ought to be quite useful if your current problem happens to be that language is being used to simulate a more abstract concept.
When I finally gave in and tried out a popular chatbot to see if it could unpack a chemical reaction description, I was initially quite stunned by how well it performed: I will admit that I was expecting to have to add an additional layer of domain specific training to get any real value, but it seems like having ingested all of the world’s patents means that that step has already been carried out. I simply asked the model to analyze a paragraph of text describing a chemical reaction and pull out all of the components and put them in a straightforward JSON structure, with some additional fields associated. The performance seemed amazing: it got almost everything right, and was even able to do things like unit conversions, and deconvolute some fairly elaborate descriptions.
And then I started throwing a substantial amount of data at it and observing trends. My initial experiment was to see if it could re-examine the 2016 patent extraction data from Daniel Lowe, and maybe leverage the incredible innovations in text extraction that have become available since then. For the kinds of reaction prediction models I’m building, I have a zero tolerance policy for bad data. If there was some way to screen the results so we could programmatically determine that each one is legit – we might be left with some fraction of the massive dataset. What fraction would we get, with zero human intervention, and almost perfect recall: one in ten? One in a hundred? Less?
The three problematic issues that seem hardest to deal with are:
- role classifications, especially with regard to picking the difference between a solvent used in the reaction rather than as part of the workup: if you ask an AI, almost every single reaction is done with ethyl acetate and hexanes
- reference context: documents frequently refer to materials (e.g. the product of X was used) or procedures (e.g. was prepared as described in Y) which the AI was not given
- hallucinations: the AI really likes to just make stuff up, and the worst part is that it’s really good at making up things that sound plausible
These are not necessarily unsolvable problems, but I don’t think that it’s going to be easy.
Where this project stands right now is a prototype interface, which is currently talking to a smaller open source LLM which can run on an ordinary computer. The rationale for veering away from the really huge commercial models is that the problem that I’m trying to solve is relatively simple and well aligned to the technology: it doesn’t need to do reasoning or inference, just pull out the right pieces and tag them.
The example below is a fairly concise description of a single step, toward the synthesis of tezacaftor, pasted into the prototype interface:
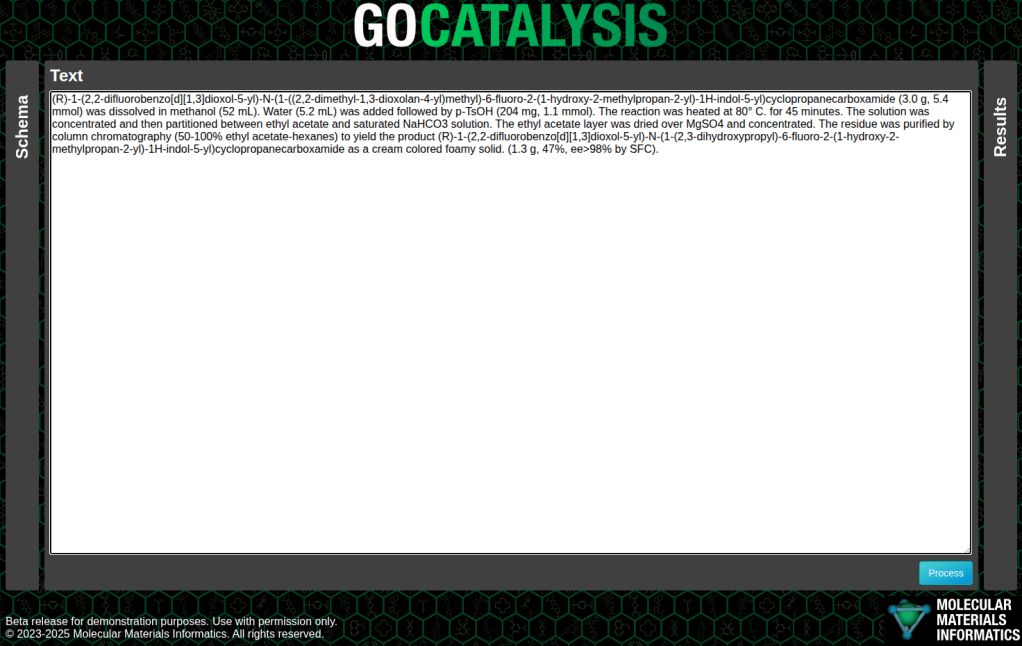

One of the reasons for making an interactive tool this early on in the exploratory phase is that I wanted to make it easy to experiment with variations on the question that is being asked of the large language model. So there’s an intermediate schema layer, which makes it easy for me to tinker with the query inputs – and under the hood, it converts the schema into the form used by the OpenAI API, which has been adopted by a whole lot of open source models.
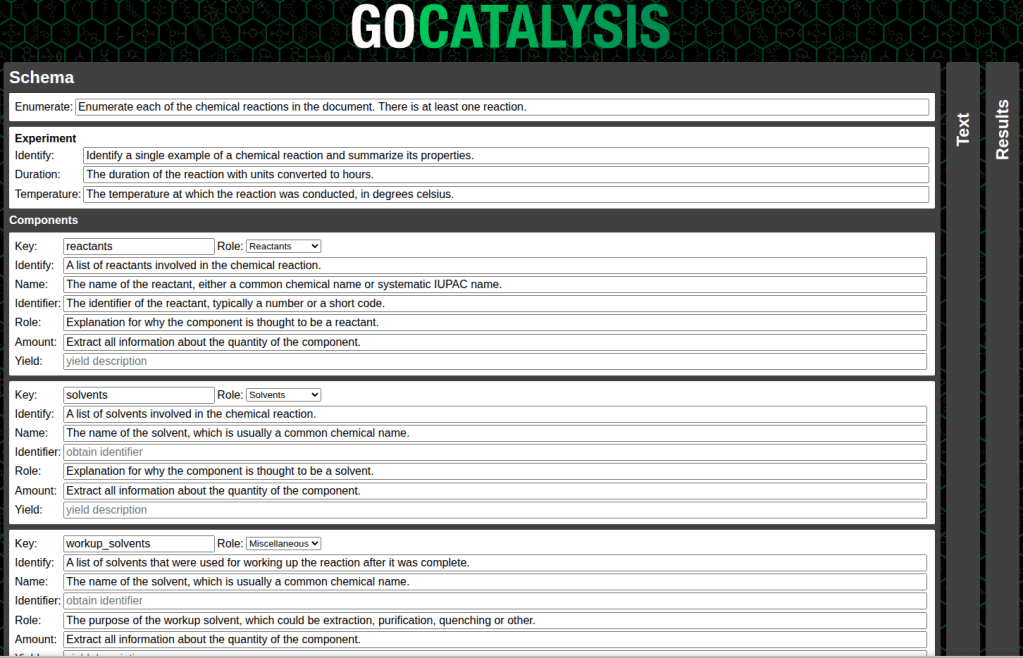

What comes back is basically a JSON hierarchy which contains a number of fields that got picked out by the LLM. That serves as a raw template for programmatic tools to take and run with:
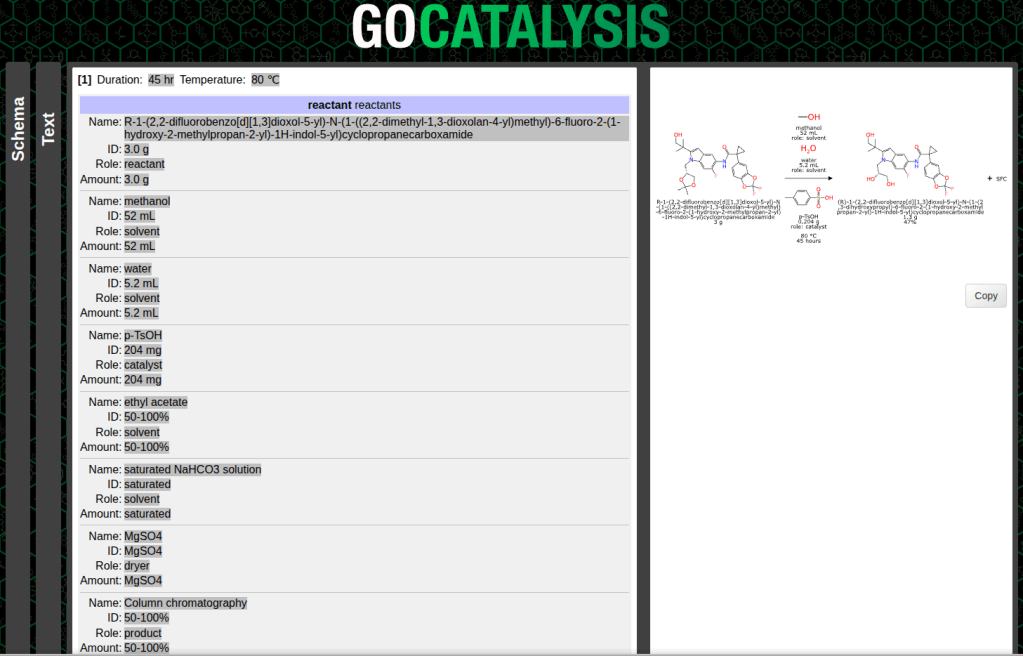
The subsequent steps involve making sense of the categories and roles, which means eliminating components that aren’t really part of the main reaction, and putting everything in its proper place: reactants, reagents, catalysts, solvents, adjuncts, products and byproducts.
Structures are converted from names, which are hopefully either IUPAC nomenclature for the reactants and products, or for the other components, something that can be cross referenced to a database of useful reaction ingredients. Quantities usually come in as mass, volume, moles or yields, depending on the role.
With structures and roles available, it is possible to perform an automated mapping, and then redepict all of the stoichiometric reactants (and then byproducts, if any) to show the common orientation:
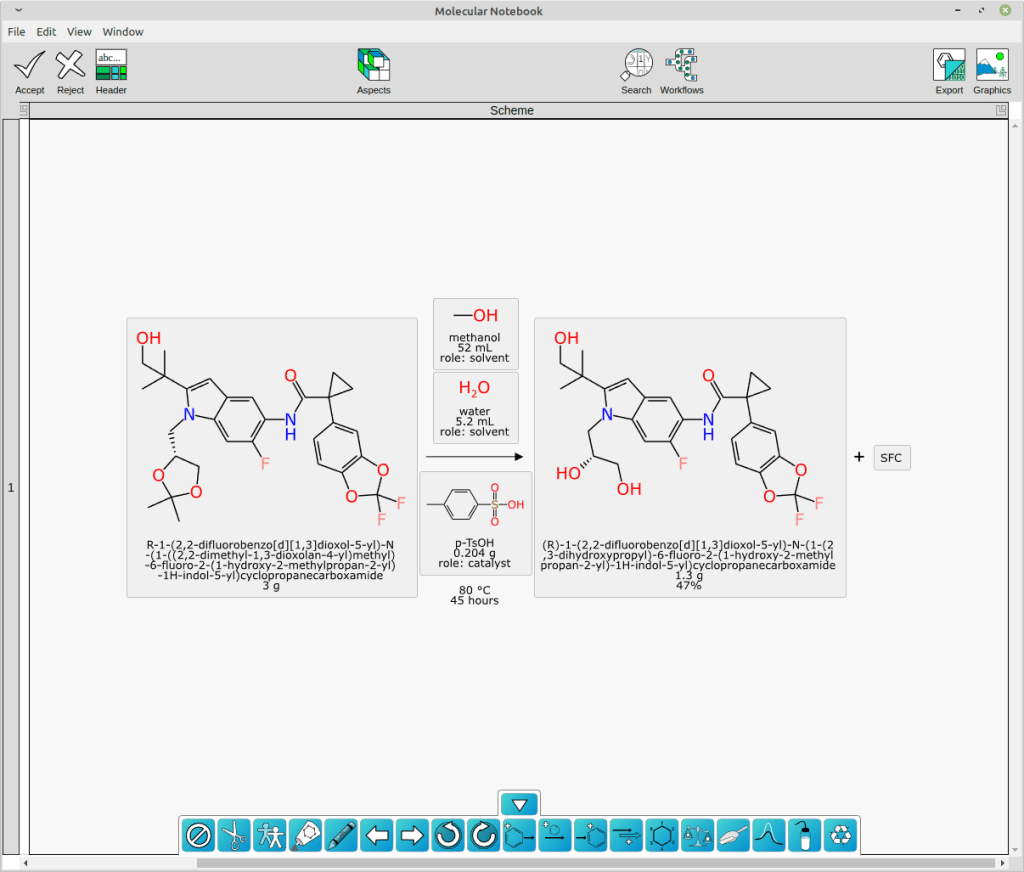
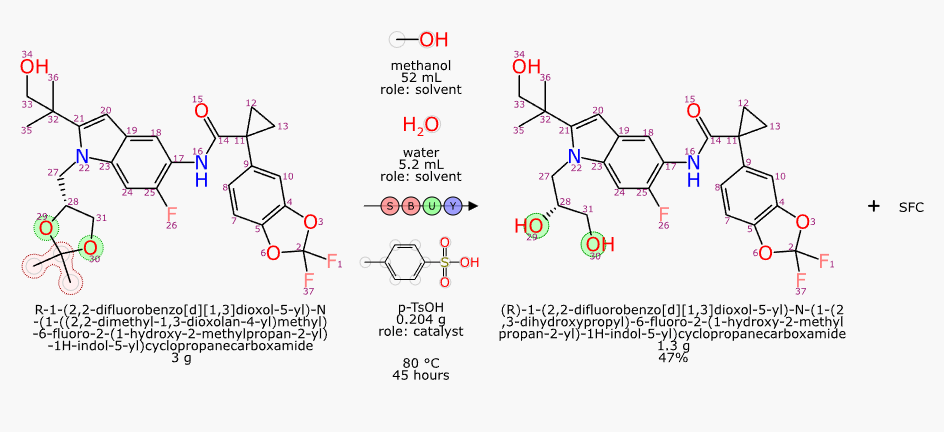
As you can see, the extraction of text content using an open source LLM followed by some domain-specific graph based deep learning models and good old fashioned cheminformatics produced a pretty good result. Not perfect, though: there is still a nonsense product named “SFC” that does not belong there. And full disclosure, I did have to add p-TsOH to my list of custom lookup names.
There is a lot more work to be done in order to refine this toolset to be useful. It is not currently obvious what workflow it would be most suited to. The idea of text mining legacy data is appealing, but the difference between a 0% error rate and a 0.1% error rate is whether or not you have to check every single record yourself. Things get harder if you want to process publications, which have a lot of contextual references inside and outside the paper, and also express a lot of content in graphical schemes and figures. The tool could alternatively be used as a very quick outlining tool, e.g. if you had a reaction description from an ELN/publication/email, you could just paste it in and get most of it right, then quickly fix up the rest.
If anyone has any thoughts on the subject, I’d certainly be interested to hear them.
