It’s been awhile since I last had the task of maintaining a chemical data warehouse using an SQL relational database. That’s not exactly a coincidence: I put in my time and did a lot of work with Oracle and MySQL back in the day, but my takeaway conclusion was that the transactional table-based systems are profoundly unsuitable for scientific data. The recently popular wave of “noSQL” database systems (such as MongoDB) are, on the other hand, quite a natural fit.
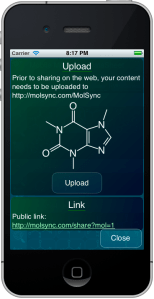
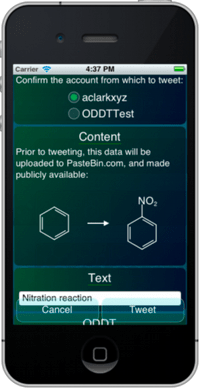
Some of the recent developments with Open Drug Discovery Teams, and content hosting for web sharing, have necessitated that the molsync.com server trade in its former stateless purity, and run a database.