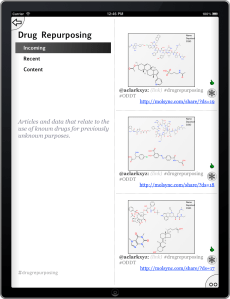
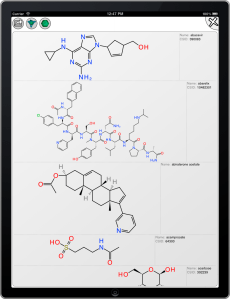
The Open Drug Discovery Teams project started out as a minimum viable product that obtained all of its data from Twitter, using a server application that periodically polls for relevant hashtags. The development version has now been extended so that topics can also be associated with RSS feeds, and each of the topics has been paired with a feed created via Google Alerts. These source documents are spliced in with the Twitter feed, e.g. the Malaria topic now contains factoids that are obtained by looking for #malaria and a timely feed of articles that Google thinks are relevant to the “Malaria” keyword.
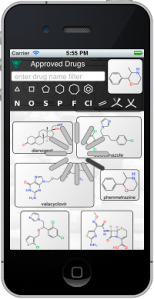
All of the crowdsourcing is still done via Twitter, regardless of where the content came from, i.e. you emit a tweet to endorse or disapprove. And using Twitter is still the appropriate way to inject content into ODDT, because you can just tweet it out and wait for it to be collected.
Addition of the Google-curated content provides a nice supplement the content stream. It will be going live soon, following a bit more testing on the development server.