Now that the molsync.com service has been upgraded so that it can produce human-readable pages with experiment details generated by the Green Lab Notebook app, it is time to demonstrate some of these. The snapshot to the right shows a screen grab of a collection of experiments that were manually keyed in from the ChemSpider Synthetic Pages service. Unlike the original data, though, the schemes have been carefully constructed so that each and every atom is accounted for in the chemical structure representations, and whenever possible all byproducts are accounted for, all stoichiometric reagents are correctly balanced, and quantities are entered in a standardised form. Continue reading
Now that the molsync.com service has been upgraded so that it can produce human-readable pages with experiment details generated by the Green Lab Notebook app, it is time to demonstrate some of these. The snapshot to the right shows a screen grab of a collection of experiments that were manually keyed in from the ChemSpider Synthetic Pages service. Unlike the original data, though, the schemes have been carefully constructed so that each and every atom is accounted for in the chemical structure representations, and whenever possible all byproducts are accounted for, all stoichiometric reagents are correctly balanced, and quantities are entered in a standardised form. Continue reading
Month: August 2014
Solvent selection in Green Lab Notebook app
 The next update of the Green Lab Notebook app brings the environmental solvent properties to the selection dialog, which is a convenient way to insert solvents into a reaction scheme. Continue reading
The next update of the Green Lab Notebook app brings the environmental solvent properties to the selection dialog, which is a convenient way to insert solvents into a reaction scheme. Continue reading
Rendering of multistep experiments for export: Microsoft Word
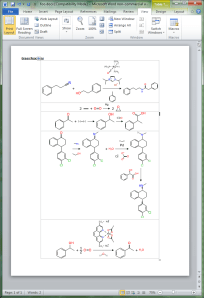
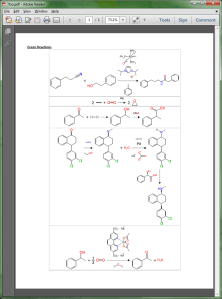
As alluded to in the previous post, one of the priorities for the Green Lab Notebook app is to make sure that it is possible to take your painstakingly drawn and cheminformatically correct reaction schemes, and produce a manuscript-quality diagram on export. This is starting to bear fruit, as the experiment-to-vector layout algorithm has been ported to the serverside (com.mmi) framework, and can now be used as the engine for creating inline graphics for tables in a Microsoft Word document:
Green Lab Notebook app now available on iTunes
 After somewhat more delay than I would have liked, the Green Lab Notebook (GLN) app is now live and available on the iTunes AppStore. The feature set that made its way into version 1.0 was crafted to ensure that editing of multistep reaction experiments and automatic calculation of green chemistry metrics are fully functional, and that importing and exporting features are complete enough to be effective. Continue reading
After somewhat more delay than I would have liked, the Green Lab Notebook (GLN) app is now live and available on the iTunes AppStore. The feature set that made its way into version 1.0 was crafted to ensure that editing of multistep reaction experiments and automatic calculation of green chemistry metrics are fully functional, and that importing and exporting features are complete enough to be effective. Continue reading
Publishing with PeerJ: first paper and first impressions
Spoiler: the experience of publishing a manuscript with PeerJ was very positive. My first paper published with them just arrived online: Fast and accurate semantic annotation of bioassays exploiting a hybrid of machine learning and user confirmation. This is a summary of the work that I was doing in San Francisco for the first half of summer 2014 (with Collaborative Drug Discovery), and it describes a hybrid machine learning/interactive method for marking up bioassay data, which is an optimised compromise between the two extremes of methods for taking plain text and turning it into semantically rich markup. Continue reading
Summer of 2014: roundup
Things have been a little quiet on this blog lately, as well as the Molecular Materials Informatics website, and indeed in the secret laboratory: and there is a perfectly good explanation for that – I’ve been away. For the last few months I’ve been quietly roosting in San Francisco, working on an interesting and exciting project at the bay area HQ of Collaborative Drug Discovery. Not wanting to put out any spoilers, but the beans will be spilled very soon via PeerJ, which should be releasing a paper with my name on it very soon indeed.
I left San Francisco a couple of weeks ago, which unfortunately means I didn’t get to attend the ACS meeting that many of you are heading towards right about now, but there is a perfectly good explanation for that, too: at the end of last month, I turned 40, and decided to celebrate this milestone by returning to my home country of New Zealand, which I have not been back to for a long time. It was a delightful trip, albeit too short: after having been away from my normal home (in Montreal) for more than 4 months, the things-to-do list was piling up quite high, as it tends to do. Continue reading