One of the habits that most chemists engage in when they sketch out structures with a pen and paper is the use of abbreviations for certain kinds of functional groups. Using abbreviations saves a lot of wasted time and ink, and it is also a useful way to draw attention to the chemistry that is relevant to the subject at hand, and away from the spinach that is coming along for the ride. For example:
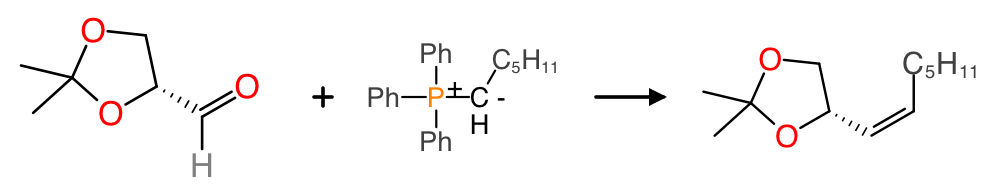 In this classical organic chemisty reaction, the phenyl groups of the Wittig reagent are abbreviated as Ph, and the unreactive alkyl chain is abbreviated as a molecular formula, because these groups are inert and not particularly exciting; the part that really matters is the reaction transform.
In this classical organic chemisty reaction, the phenyl groups of the Wittig reagent are abbreviated as Ph, and the unreactive alkyl chain is abbreviated as a molecular formula, because these groups are inert and not particularly exciting; the part that really matters is the reaction transform.
Organic chemists typically have a choice as to whether to use abbreviations as a short-hand notation, but when it comes to inorganic structures, particularly coordination complexes, there isn’t really much of a choice. For example, consider one of the compounds that I synthesised during my Ph.D. research:
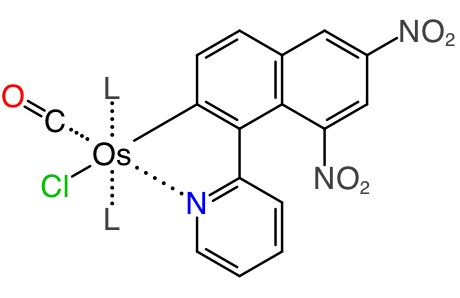 This is a clear and concise diagram, and it calls attention to the parts that matter: the coordination about the central metal, and the structure of the organometallic ligand, which is the point of the research. Now, if this same structure were to be drawn in a way that is complete from a cheminformatics point of view, it would need to include all of the heavy atoms, which would mean the best way to draw it would be more like this:
This is a clear and concise diagram, and it calls attention to the parts that matter: the coordination about the central metal, and the structure of the organometallic ligand, which is the point of the research. Now, if this same structure were to be drawn in a way that is complete from a cheminformatics point of view, it would need to include all of the heavy atoms, which would mean the best way to draw it would be more like this:
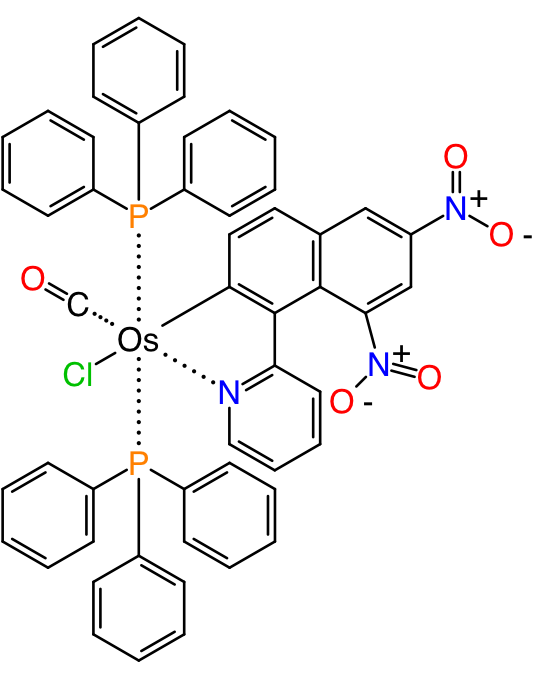 There is probably not much need to go into why the first rendition is a better way to communicate information from one chemist to another. When there is a computational algorithm involved, or in fact a fellow chemist who is not fully briefed on context, such as the implicit understanding that L is a placeholder for triphenyl phosphine, then additional information has to be added to ensure that the real intent of the structure is not lost.
There is probably not much need to go into why the first rendition is a better way to communicate information from one chemist to another. When there is a computational algorithm involved, or in fact a fellow chemist who is not fully briefed on context, such as the implicit understanding that L is a placeholder for triphenyl phosphine, then additional information has to be added to ensure that the real intent of the structure is not lost.
This example is hardly the most direly in need of an abbreviation system – it gets much worse. A quick look through Cotton & Wilkinson will confirm that being able to represent inorganic structures legibly most of the time is going to need an alternative to an all-heavy-atom visual representation.
One approach to making use of abbreviations without losing information is to have a centralised repository of standard monikers, so users can type on whatever they want for atom labels, and as long as it is on the official list, software can expand it out. Apart from the obvious complications involved in maintaining such a list, this is a nonstarter because abbreviations have never been standardised, and never will be. While some common ones, such as Ph, are more or less universally recognised, others such as L are used for all kinds of purposes. Standardisation is a relative term: groups of chemists often have a shared understanding of what certain abbreviation symbols mean, but another group may not know, or use it for something different.
The other alternative is to store the definition of the abbreviation within the structure itself, which removes all of the really intractible problems, swapping them for some that are merely tricky.
At the moment, the Mobile Molecular DataSheet (iOS version) is undergoing a retrofit to add an abbreviation system. It is nominally complete, but since this feature impacts seemingly unrelated functionality within the app, version 1.2.6 is undergoing a longer test cycle than usual. Which is why this feature is being written up in a blog about what’s coming up soon, rather than an article about what’s already there.
In the new version, any atom, or terminal fragment, can be converted into an inline abbreviation. After the conversion, one atom remains, as a placeholder. Stored behind the scenes, in the atom-expansion fields, is the actual structure of the abbreviation fragment (hence the term “inline”).
The following pictures show some of the user interface concepts:
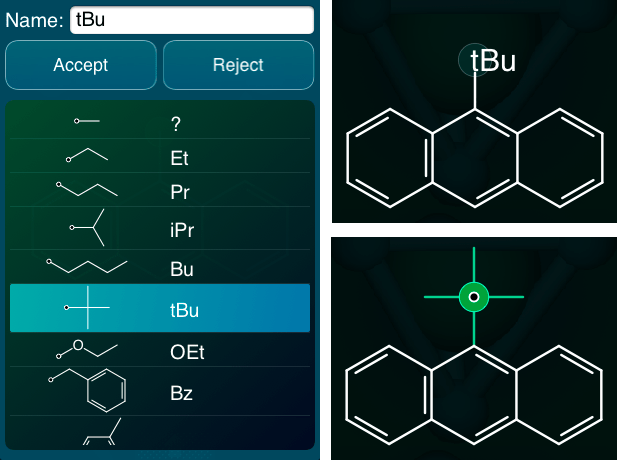
The dialog-style panel on the left shows the process of selecting an abbreviation, either for creating a new one or changing the definition of an existing one. The list is built from the template datasheets, and any abbreviations that have been defined in the current molecule, or other molecules in the current datasheet.
The structure is normally displayed as shown on the top right, i.e. the label of one of the “atoms” is tBu. When an abbreviation placeholder is the current atom, a full expansion of the abbreviation is shown in green, which is demonstrated on the bottom right, so from within the atom editor, it is easy to keep track of which atoms are abbreviations and what they stand for.
The abbreviation placeholder atom must be strictly terminal, i.e. you can’t have an abbreviation that is joined to more than one thing, but within the abbreviation fragment itself, there can be multiple connection points. This means that multidentate connections can be collapsed into inline fragments, e.g. the cyclopentadienyl ligand:
On the left is the typical display, and on the right is what is shown when the abbreviation is the current atom.
Because of the way abbreviations are defined and stored, it is always possible to use a simple algorithm to expand out the full all-heavy-atom structure, which means that it is possible to use abbreviations and be sure that derived properties will not be broken: for example, molecular weight and formula calculations are done correctly. The formula for the above structure is C7H5ClFeO2, not C2CpClFeO2!
There is a bit more to this feature than the examples shown above, but that is enough to get an idea of what is coming up for MMDS v1.2.6.

A timely example of the kinds of problems this is supposed to circumvent:
http://www.chemconnector.com/2011/04/29/markush-misrepresentations-in-chemspider/