 Until earlier this week, the public facing instance of the BioAssay Express was basically read only: we carried out our curation exercises by passing files around. As of now, though, it is possible to authenticate yourself using ORCID and submit your annotations. From that point your changes will be stored in a special holding bay until they are approved. This feature is enabled on the beta preview version.
Until earlier this week, the public facing instance of the BioAssay Express was basically read only: we carried out our curation exercises by passing files around. As of now, though, it is possible to authenticate yourself using ORCID and submit your annotations. From that point your changes will be stored in a special holding bay until they are approved. This feature is enabled on the beta preview version.
The BioAssay Express project has been on the open internet for a number of months now. One of the reasons why we were able to get away with posting our work in public for everyone to access is that we cut an important corner: the project had no way to authenticate users, and no way to submit content, which means that a significant amount of energy could be diverted away from worrying about security, and channelled directly into getting the science & the user experience right.
That limitation has now been lifted, though we did take the path of least resistance: the authentication process piggybacks on top of the ORCID service. ORCID stands for Open Researcher & Contributor ID, and the objective of that project is to provide a way for scientists to identify themselves unambiguously. If you wonder why that is necessary, use some service like Google Scholar and type in your name, then find out about all of the weird & wonderful research that you don’t remember doing. The ORCID project provides an OAuth service, which is nerdspeak for when a website temporarily redirects you to a service that you’re already signed up for in order to confirm your identity. This is done as an alternative to making you sign up for another service, so you don’t have to remember yet another password.
The front page of the beta version of the BioAssay Express now looks like this:
Clicking on the Login button brings up a dialog that is filled with ORCID content. You will be prompted to make it known that you are you, or create a new ID if you don’t have one:
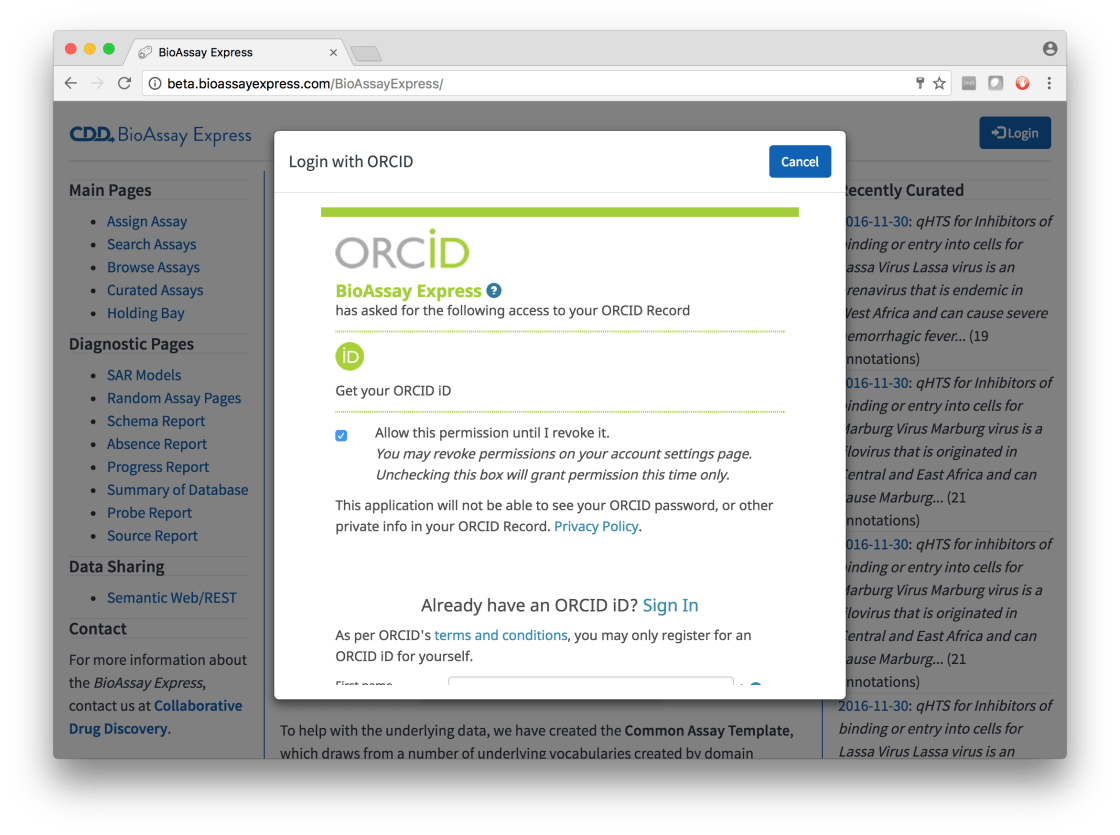
You must then confirm that you are OK with letting the BioAssay Express know a little bit about you. This information is basically your identifier, name and email address. Assuming that this isn’t too much of an invasion of your privacy, continuing on will make the Login button change to Logout, with a mouseover that mentions who the system currently thinks you are:

As if all this wasn’t exciting enough, once you are logged in, there is currently one thing that you can do that you couldn’t do before: Submit annotations for an assay:
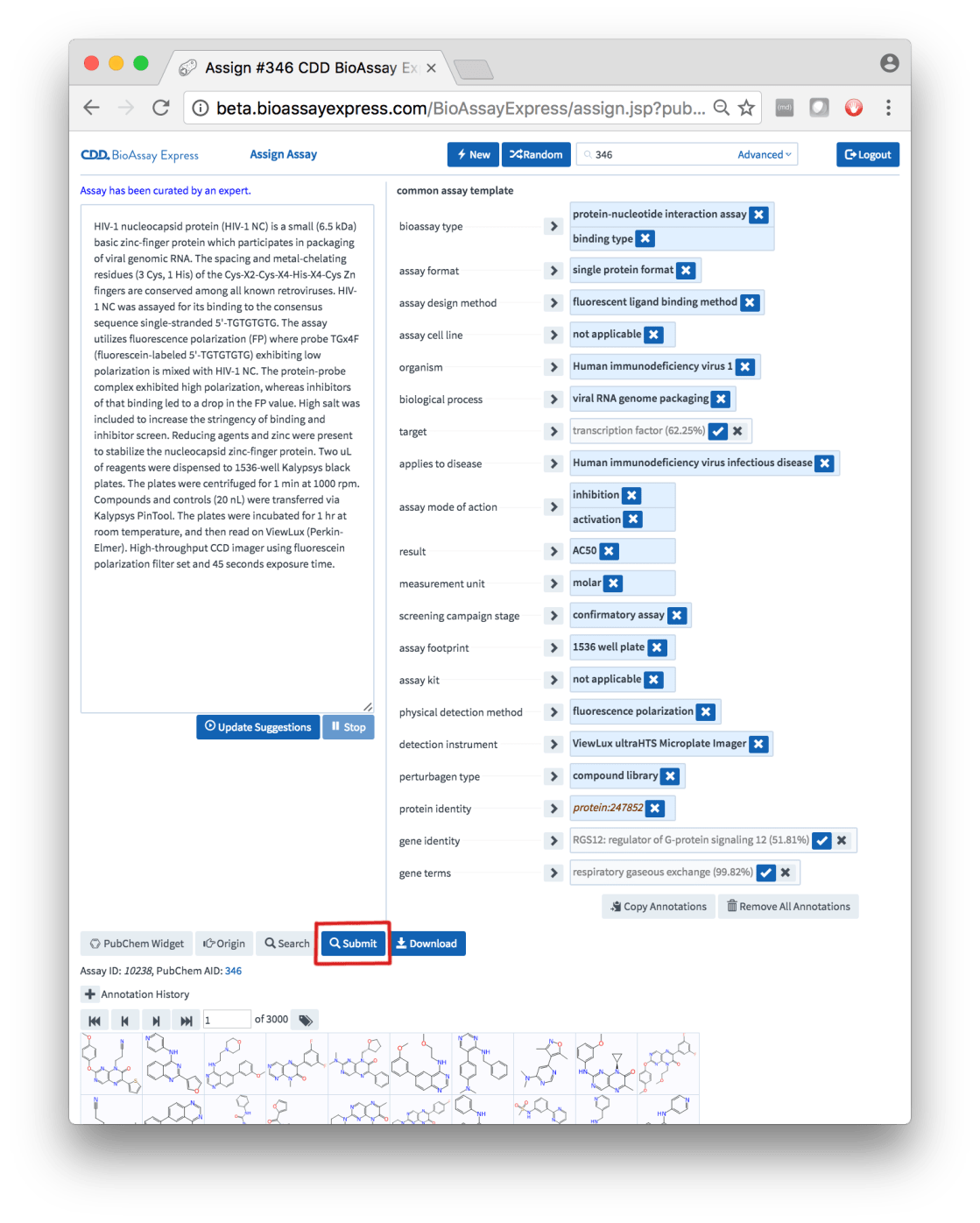
Near the bottom of the snapshot shown above is the newly added button. Up until this point it was possible to annotate a variety of assays that had been preselected from PubChem, or annotate new assays of your own, but the only deliverable was via the Download button, which allows you to acquire a file containing your hard work – with the caveat that you are in charge of what happens to it from that point onward. The Submit button is closer to what you would expect from a functional web service: the changes are stored on the server itself, and you are free to move onto your next task.
Immediately prior to this new functionality, the BioAssay Express was amended so that it now stores a record of all modifications: the history of additions and deletions can be viewed easily on the assignment page. Similarly, any submissions that are sitting around in the holding bay can also be viewed while awaiting approval. This means that if you “change” a bunch of content, even if the administrator (currently me) is a bit lax about what gets approved, there will still be a clear record of what was there before.
This new authentication/submission capability is an important step for the project’s journey toward becoming an enterprise ready product, but it also has a more immediate implication. The baseline content for the publicly facing service has been pulled out of PubChem, and followed by our internal efforts that involved annotating 3500 of them. That’s a lot of assays, but there are a lot more waiting to be marked up – even if we only consider the assays that have full text descriptions. This means that if you – dear reader – see an assay that happens to be one of yours, or you know it well enough to select some or all of the appropriate descriptive terms, then you are not only welcome but also empowered to do exactly that!