The open source project SketchEl, hosted by SourceForge, is a cheminformatics-focused structure editor that I wrote back at the dawn of time (also known as the year 2005). The latest version, 1.56, adds in MDL MOL extensions that allow this popular legacy format to be used for reading and writing structures that use zero-order bonds, and non-computable virtual hydrogens, as described in J. Chem. Inf. Model., 51, 3149-3157 (2011).
The open source project SketchEl, hosted by SourceForge, is a cheminformatics-focused structure editor that I wrote back at the dawn of time (also known as the year 2005). The latest version, 1.56, adds in MDL MOL extensions that allow this popular legacy format to be used for reading and writing structures that use zero-order bonds, and non-computable virtual hydrogens, as described in J. Chem. Inf. Model., 51, 3149-3157 (2011).
It’s written in pure Java, uses the Swing UI toolkit, and has a fairly powerful (albeit uneven) feature set. The motive for writing it was quite simple, and rather common for open source projects: I needed it. At the time the list of structure editors that ran on Linux was limited to a handful, and they were crude and/or broken, whereas there were a number of free (“as in beer”) editors for Windows that were adequate. I needed to test software that involved wrangling 2D structure representations, and having to get out of my chair and walk over to a Windows computer to view a structure got old pretty fast.
Because the SketchEl program was designed for cheminformatics, it is heavily biased in favour of maintaining a datastructure that captures the scientific meaning of a chemical structure, and in turn it eschews any kind of recorded property that does not have a well defined meaning. This lead to the invention of the SketchEl molecule format, which is used by all products from Molecular Materials Informatics, because it is extremely lean, minimalistic, well defined, extensible, forward- and backward-compatible, and designed to represent the chemistry of molecular entities from throughout the periodic table, not just the myopic subcategory of organic druglike molecules.
A bit more than a year ago, I published a paper (mentioned above), entitled “Accurate Specification of Molecular Structures: The Case for Zero-Order Bonds and Explicit Hydrogen Counting“. The SketchEl molecule format is roughly equivalent to MDL MOL for most purposes, except that it allows bonds with order 0, and explicit specification of virtual hydrogen counts. These two seemingly innocuous and trivial features make it possible to fix a vast range of failures in chemical databases, if used properly. But of course as everyone knows, convincing a whole industry to use a different standard data format tends to be rather Quixotic, so it’s a lot more realistic to try to encourage people to adapt their software to extend the incumbent format.
Extending the legacy format isn’t just a matter of adding some extra fields: it’s also very useful to try to downgrade the structure, to the most reasonable option for legacy software, then use the extension fields to re-upgrade the structure back to what it ought to be, for more enlightened software.
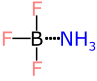
For example, consider the Lewis acid:base adduct BF3:NH3. The right way and the proper way to draw this is:
Note the dotted line between boron and nitrogen (“zero-order”), which is used to indicate that it is not a covalent bond, and therefore should not be included in the valence calculation. Thus the nitrogen atom has no attached single bonds, and therefore needs to be topped up with 3 extra implicit hydrogen atoms to make up its Lewis octet shell. So far so good.
Now if the structure were exported to a legacy format, like MDL MOL, one approach is to convert the zero-order bond to single (because 0 is not permitted), then use additional fields to indicate to modern software that the bond is, in fact, a zero-order bond. This is easy to do with MDL MOL, so it the file would look like this:
SketchEl molfile 5 4 0 0 0 0 0 0 0 0999 V2000 -3.5500 3.8000 0.0000 B 0 0 0 0 0 0 0 0 0 0 0 0 -2.0500 3.8000 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 -3.5500 5.3000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 -5.0500 3.8000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 -3.5500 2.3000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 1 2 1 0 0 0 0 1 3 1 0 0 0 0 1 4 1 0 0 0 0 1 5 1 0 0 0 0 M ZBO 1 1 0 M END
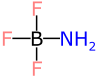
The first bond (B-N) is listed as a single bond, then the ZBO modifer extension is added, in order to allow modern software to adjust the bond order to zero. That’s fine for modern software, but legacy software that does not understand ZBO will ignore it, and so will parse out the structure as:
This is not just a suboptimal description of the molecule, it’s just flat out wrong, and not in a pedantic way. The boron atom has the wrong valence, which means that some software might be tempted to fix that by giving it a negative charge. The nitrogen is assigned the wrong number of implicit hydrogens, hence the molecular formula is wrong. In the worst case scenario, this flaw could make your experiment not work, or your flask explode, or at the very least prevent you from finding the entry in a chemical database.
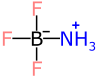
As it happens, there is a reasonable way to represent this structure using only the fields that are available to the legacy MDL MOL format:
It’s not ideal. It does not do a very good job of describing the chemistry of the bonds or the environment of the atoms, but it does at least lead to the right valence, and the correct molecular formula. That means if you’re allowing an electronic lab book to calculate your quantities for you, it will at least tell you the correct amount to add.
Not all nonorganic bonds have this kind of “reasonable hack”, but quite a few of them do, and it should be exploited. So the ideal way to write out this structure in a legacy format is to include both representations, so that (1) legacy software gets the charge-separated version that it can do some useful work with, and (2) modern software can parse the proper version.
The extensions are written up like this:
SketchEl molfile 5 4 0 0 0 0 0 0 0 0999 V2000 -3.5500 3.8000 0.0000 B 0 5 0 0 0 0 0 0 0 0 0 0 -2.0500 3.8000 0.0000 N 0 3 0 0 0 0 0 0 0 0 0 0 -3.5500 5.3000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 -5.0500 3.8000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 -3.5500 2.3000 0.0000 F 0 0 0 0 0 0 0 0 0 0 0 0 1 2 1 0 0 0 0 1 3 1 0 0 0 0 1 4 1 0 0 0 0 1 5 1 0 0 0 0 M CHG 2 1 -1 2 1 M ZCH 2 1 0 2 0 M ZBO 1 1 0 M END
Note that the official MDL MOL parts – the atom and bond blocks, and the CHG extension, encode the charge separated version with a B-N single bond. The ZCH and ZBO non-standard extensions, which are only read by modern software, overwrite the charge and bond order for the affected atoms.
Long story short, the MDL MOL extensions and the charge separation algorithm necessary to produce this output have both been added to the SketchEl package, so it is possible to draw inorganic compounds, save them as .mol files, read them back into SketchEl (or other compatible products such as the Mobile Molecular DataSheet), and also feed them to other software that doesn’t understand the enhanced fields, and expect the results to be about as good as they could be, under the circumstances.
The SketchEl program is released under the Gnu Public License, which means that you’re welcome to grab the source code and incorporate that into your own GPL software. Or if you just want to look at the algorithms, and the extensions, then reimplement it in a proprietary package, you’re equally welcome to do that.