The Royal Society’s Medicinal Chemistry Toolkit app has been up on the AppStore since late last year, but a couple days ago it got updated with some major new functionality: an interactive tool that allows a structure to be drawn, and various properties to be calculated from it. If you can’t guess who supplied the sketcher and the structure-derived calculations, you’re probably not a regular on this blog.
We have been working together on this since last year. The embedded sketcher component is provided by a library that is closely related to the MMDSLib toolkit that powers a variety of apps, either from Molecular Materials Informatics directly, or as a component of third party apps. This particular incarnation adds some features that were built or tuned especially for the occasion. As well as being able to calculate simple properties like molecular formula & weight, the implementation of Crippen’s log P algorithm was ported to mobile, and also carefully validated (which is harder than it sounds, in the absence of an actual validation set). This has been available by my implementation via webservice for some time, but in order to make it work directly on an app – without sending anything out over the internet – a significant amount of functionality had to be ported first, namely substructure searching, and all of the algorithms that it is based upon.
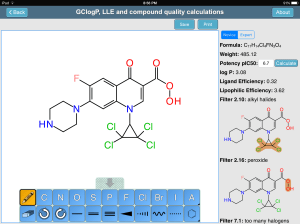
After that, the next step was to implement the published Astra-Zeneca filters within the app, in order to automatically highlight certain kinds of structures that have been established to be bad ideas that can be safely triaged from collections of potential drug candidates. While implementing these filters seemed like a moderately complex task, given that the substructure searching capabilities had already been migrated to the iOS (Objective-C) platform, it ended up being rather demanding: the filters have evolved a bit within Astra-Zeneca since the publication of the paper, and after working with the original team, it turns out that some of the queries are a little more complex than the summary descriptions would suggest. Before this task was done, the mobile substructure search ended up supporting substructure sub-queries, with infinite recursion and logical clauses. Incidently, these improvements are what made it possible to also implement the PAINS filters within mobile apps. Many passes over the ChEMBL dataset later, and many midnight sessions poring over lists of pass/fail validation case studies, it was agreed that the algorithm used by the Medicinal Chemistry Toolkit app is indeed a good implementation of what is being used for real world drug discovery at Astra-Zeneca.
Because these filters are sort-of proprietary, that means that this particular app is the only place you can find them in a publicly available product. Oh, and it’s free, as long as you have an iPad.