 Abbreviations are a common shorthand used by chemists to alleviate the tedium of drawing out structures by hand, and in many areas – including much of organometallic and inorganic chemistry – use of abbreviations is more or less essential to achieve any kind of visual clarity. This presents a challenge for cheminformatics, though, because there is no universal dictionary for abbreviations: different research groups use different shorthand notations for their own nefarious purposes. Now the beta version of XMDS is catching up to MMDS by adding the ability to conveniently define inline abbreviations, which are readable to humans in diagram form, but also completely well defined behind the scenes for the benefit of machines.
Abbreviations are a common shorthand used by chemists to alleviate the tedium of drawing out structures by hand, and in many areas – including much of organometallic and inorganic chemistry – use of abbreviations is more or less essential to achieve any kind of visual clarity. This presents a challenge for cheminformatics, though, because there is no universal dictionary for abbreviations: different research groups use different shorthand notations for their own nefarious purposes. Now the beta version of XMDS is catching up to MMDS by adding the ability to conveniently define inline abbreviations, which are readable to humans in diagram form, but also completely well defined behind the scenes for the benefit of machines.
As was expounded upon recently in an article provocatively entitled “Machines first, humans second: on the importance of algorithmic interpretation of open chemistry data” (Journal of Cheminformatics 2015: open access), the blatant failure of many molecular description practices to encode all of the atoms in the molecule (i.e. make it possible to derive the molecular formula correctly) is an embarassment to our profession. For the traditional domain of cheminformatics, which implicitly restricts itself to the realm of organic compounds with banal Lewis-compliant bonding, the best practice recommendation for use of most software is to simply avoid using abbreviations at all. Unfortunately there are a great many molecules out there in the wild with atoms that have names like “Et”, “iPr”, “Ph”, etc., and there is a common expectation that the smarter cheminformatics algorithms ought to be able to interpret these adequately well.
Like most ad hoc conventions that have accumulated over decades, there are areas of common ground that everyone agrees on: it is unlikely you will find a chemist who will argue that “Et” means anything other than ethyl (C2H5), but try asking an inorganic chemist what “L” stands for, without any context. Obviously it stands for the ligand – that much is clear – and one might go further to say that it is the ligand that that particular chemist uses the most, and considers least relevant to the actual chemistry of interest. In the middle of these two extremes are any number of conventional variations (e.g. But vs tBu) and abbreviations that are unique to one particular group of chemists who became tired of drawing out some common fragment. And then it gets weird when quasi-queries show up (like “R” or “X” or “Z”).
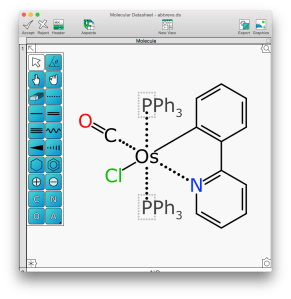
The point is that having a dictionary of abbreviations is not the solution to the problem. The solution is to use an inline encoding of the abbreviation, which is defined neatly as an extension to the minimalistic SketchEl molecule format. While the Mobile Molecular DataSheet (MMDS) app has had this feature for awhile, it is now coming along nicely in the OS X Molecular DataSheet (XMDS) desktop app. For example, the structure shown above – which is one of the new compounds I made back in my graduate student days – the two triphenylphosphine (PPh3) abbreviations do something interesting when the mouse is moved over either of them:
The abbreviation placeholder atom is replaced by its full definition as a “ghost fragment” in a lighter shade of grey. As you can see, it’s rather overbearing: if the two phosphine ligands were drawn out completely, first of all the geometry would be awkward and unpleasing to the eye, and secondly it would dominate the structure. For communication-to-human purposes this is suboptimal, because these large ligands are actually really boring. The reason they are there is basically to act as umbrellas to protect the sensitive areas; the interesting chemistry is happening at the metal centre.
The way this is done without compromising any of the cheminformatics (i.e. the molecular formula is easily derivable, the bond orders still add up with the right valences and oxidation states, etc.) is by encoding the fragment within the definition of the atom itself.
Creating a new inline abbreviation with the XMDS desktop app can be done in several ways. If the abbreviation of interest happens to be included in the list of templates, it can be grafted on in abbreviation form with a couple of clicks. For getting a bit more creative, the way to go is to draw out the entire abbreviation in full (with optional sub-abbreviations), select the terminal fragment, then hit one of the make-abbreviation buttons:
In this sequence, the 3rd nitro group has been drawn out in full, and selected. Hitting the “abbreviate group” command subsumes the fragment, then brings up the dialog for renaming the atom placeholder, which in this case is manually entered as “N|O{3}“, which is rubric for NO2 (forward) or O2N (backward). An alternate command button – “abbreviate formula” – does the same thing, except that the label is automatically labelled as the molecular formula, which is a great time saver in some cases. Abbreviations can be removed, or expanded out permanently with a single button press.
One of the things about abbreviations is that you probably don’t want to have to draw them manually any more often than necessary. For the common abbreviations that feature in the prepackaged templates, this is easy enough to solve: the existing abbreviations are compiled into a list at runtime, and can be selected from a list:
Which is all very well, but if you were to define an abbreviation called “Foo“, for a series of molecules that you don’t feel like sketching out in full detail every time, the idea is to draw the fragment out in its full atomic glory once and convert it into an abbreviation. One solution is to copy the Foo atom placeholder onto the clipboard and paste it back in when necessary – but that isn’t always the most convenient. To make it as accessible as possible, each time XMDS opens a new datasheet, it scans through looking for any inline abbreviations in any of the molecules, and scrapes anything it finds, adding them to the temporarily global collection. These will now appear in the list of available things.
In summary, inline abbreviations are actually really important for many kinds of cheminformatics, especially non-organics, and regardless of your domain, you need to be careful to either do it right or not do it at all. XMDS, like MMDS before it, is being designed to make doing it right as painless as possible.




