Some years ago I bought a domain name on a whim – orangometallics.com. To those of us who have dwelled within a certain chemistry subdiscipline, this is immediately obvious as a comedically simian misspelling of organometallics. Unlike the proper term, the domain name was available at a very modest price, so I decided that my company needed to own that.
Continue readingMolecule rendering: the small things
And also happy 2024 to everyone, I haven’t been writing much lately on account of juggling work and family life. The topic of this post is two aesthetic molecule drawing improvements that could be described as highly unfavourable from the point of view of the effort:reward ratio, but sometimes these things just bother you for long enough that they bubble up to the top of the to-do list.
Continue readingBioAssay Express is now open source
11 September 2023
The BioAssay Express is being released as an open source project, under the Apache 2.0 license. The short description of this license is that it is permissive, and essentially the only restriction is acknowledgment.
BioAssay Express is a grant-funded project to bring semantic web annotations to bioassay protocols, using vocabularies such as the BioAssay Ontology (BAO) to enrich descriptions that are primarily stored as text. Because of the universality of ontology terms, this means that annotated assays take on standardized meaning and can be processed by machines as effectively as they can be understood by scientists. This is a canonical example of the application of FAIR data principles (F = Findable, A = Accessible, I = Interoperable, R = Reusable).
Continue readingWebMolKit: now with Apache for all
WebMolKit is a cheminformatics library that I’ve been working on for a long time: it runs on all kinds of JavaScript engines (browsers, desktop via Electron, command line via NodeJS). Its flagship feature is a powerful chemical sketcher, but it also has many supporting functions for handling molecules. As of now, the licensing terms have been switched to Apache 2.0, which basically means you are allowed to use it for non-open projects, as long as proper credit is given.
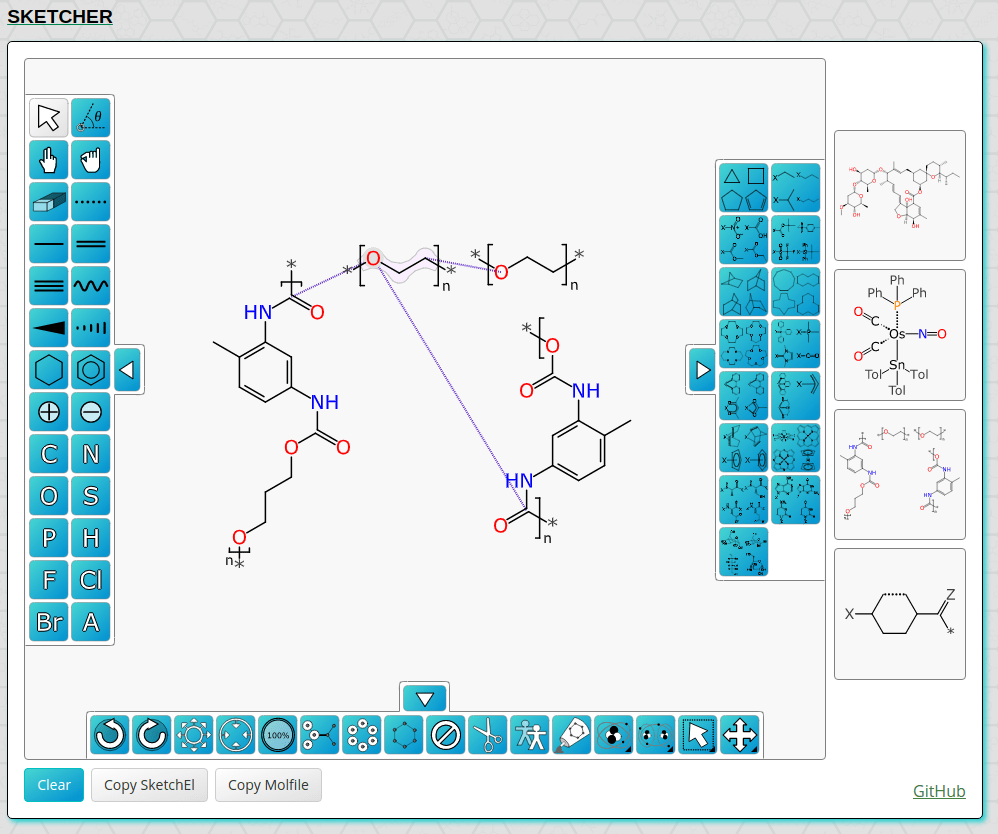
WebMolKit Sketcher
After much procrastination, there’s an updated demo of the WebMolKit sketcher on the main home page for Molecular Materials Informatics. The sketcher is open source, runs on any web platform, and has some fairly powerful functionality built into it.
Continue readingCall for papers: cheminformatics workflows
The Journal of Cheminformatics is organising a Special Collection entitled “Biomedical Data Analyses Facilitated by Open Cheminformatics Workflows” which encourages researchers to publish their workflows for gathering, preparing, curating and cleaning data. This resonates well with a growing explicit awareness within the community that data quality isn’t just an important thing, it’s the important thing.
Continue readingMixtures: a collection of online presentations
In the spring of 2021, writing from the comfort of my home office, it’s hard to even remember what it’s like to pack my bags and head out to stand in front of a room and tell people about what I’ve been doing lately. The delights of airport security, jetlag, hotel wifi and bad coffee are not exactly missed, but pretty much everything else is. Results do still get communicated, though, and one nice thing about delivering a webinar is that you can be pretty sure it will be recorded for the benefit of the whole internet for all time (or as may be the case, some of the time).
Continue readingCoordination InChI for inorganics: now with stereochemistry
About a year ago I described the results of a preliminary project to scope out the possibility of making the InChI identifier play nice with inorganic & organometallic complexes. There’s now a followup increment that does likewise for some of the higher valence stereochemistry centres that are found in these complexes.
Continue readingOnline mixtures demo, with MInChI generator
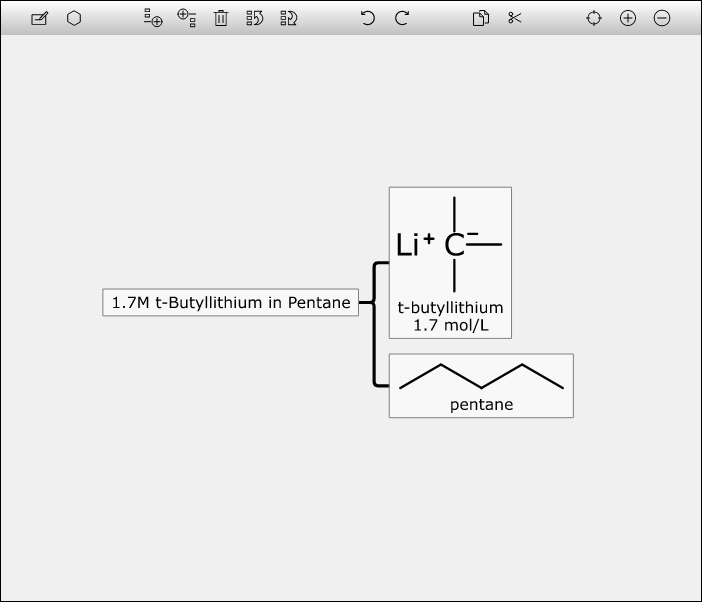 Drawing chemical mixtures can be done online, with a conversion feature to generate Mixtures InChI (MInChI) notation. Pseudomixtures from Molfiles are now enumerated automatically when pasted in. The tools for working with machine readable mixtures, using the web platform, are open source. Continue reading
Drawing chemical mixtures can be done online, with a conversion feature to generate Mixtures InChI (MInChI) notation. Pseudomixtures from Molfiles are now enumerated automatically when pasted in. The tools for working with machine readable mixtures, using the web platform, are open source. Continue reading
Adding in anti-viral model for COVID-19 predictions
 A newly available collection of antiviral structures from Chemical Abstracts has been made available, and has now been shoehorned into a model that can be used online to evaluate potential antiviral drug candidates for COVID-19. The tool can be found at https://molmatinf.com/covid19. Continue reading
A newly available collection of antiviral structures from Chemical Abstracts has been made available, and has now been shoehorned into a model that can be used online to evaluate potential antiviral drug candidates for COVID-19. The tool can be found at https://molmatinf.com/covid19. Continue reading