The last update of the MolSync site added a visual overhaul and molecule searching. Today’s update brings along the basic UI for reaction searching, and preliminary functionality for various kinds of reaction searching by structure. Continue reading
The last update of the MolSync site added a visual overhaul and molecule searching. Today’s update brings along the basic UI for reaction searching, and preliminary functionality for various kinds of reaction searching by structure. Continue reading
molsync
MolSync web structure searching
 As mentioned in the previous post, the MolSync (.com) website and the technology behind it have been moving forward rapidly. The public-facing deployment now shows a proof of concept page for performing molecule searches: molsync.com/search/molecule.php. Continue reading
As mentioned in the previous post, the MolSync (.com) website and the technology behind it have been moving forward rapidly. The public-facing deployment now shows a proof of concept page for performing molecule searches: molsync.com/search/molecule.php. Continue reading
MolSync overhaul: back to the web, now with reactions too
 Things have been a bit quiet in these parts lately, but not due to inactivity: far from it. In between working on some exciting projects with Collaborative Drug Discovery, I have been quietly making rapid progress on several important key technologies. These include the OS X Molecular DataSheet (XMDS), presiding over a growing collection of reaction data, and most recently a complete overhaul of the MolSync website, which provides cheminformatics support services of various kinds. Continue reading
Things have been a bit quiet in these parts lately, but not due to inactivity: far from it. In between working on some exciting projects with Collaborative Drug Discovery, I have been quietly making rapid progress on several important key technologies. These include the OS X Molecular DataSheet (XMDS), presiding over a growing collection of reaction data, and most recently a complete overhaul of the MolSync website, which provides cheminformatics support services of various kinds. Continue reading
A rant about data quality: machines first, humans second…
 Recently one of my papers emerged through the publication system of Journal of Cheminformatics, entitled “Machines first, humans second: on the importance of algorithmic interpretation of open chemistry data“, co-authored with Antony Williams and Sean Ekins, and incorporated into the JC Bradley Memorial Issue. Spoiler alert: the paper is about how if you’re publishing open lab notebook data without adhering to rigorously defined standards for machine readability, then you’re mostly wasting your time, and arguably making the open data situation even worse than it already is. The tone of the article is a bit less polite than I normally try to be, so fair warning, but it’s all for a good cause.
Recently one of my papers emerged through the publication system of Journal of Cheminformatics, entitled “Machines first, humans second: on the importance of algorithmic interpretation of open chemistry data“, co-authored with Antony Williams and Sean Ekins, and incorporated into the JC Bradley Memorial Issue. Spoiler alert: the paper is about how if you’re publishing open lab notebook data without adhering to rigorously defined standards for machine readability, then you’re mostly wasting your time, and arguably making the open data situation even worse than it already is. The tone of the article is a bit less polite than I normally try to be, so fair warning, but it’s all for a good cause.
Sharing experiments: using ChemSpider Synthetic Pages examples and a trip down memory lane
 Now that the molsync.com service has been upgraded so that it can produce human-readable pages with experiment details generated by the Green Lab Notebook app, it is time to demonstrate some of these. The snapshot to the right shows a screen grab of a collection of experiments that were manually keyed in from the ChemSpider Synthetic Pages service. Unlike the original data, though, the schemes have been carefully constructed so that each and every atom is accounted for in the chemical structure representations, and whenever possible all byproducts are accounted for, all stoichiometric reagents are correctly balanced, and quantities are entered in a standardised form. Continue reading
Now that the molsync.com service has been upgraded so that it can produce human-readable pages with experiment details generated by the Green Lab Notebook app, it is time to demonstrate some of these. The snapshot to the right shows a screen grab of a collection of experiments that were manually keyed in from the ChemSpider Synthetic Pages service. Unlike the original data, though, the schemes have been carefully constructed so that each and every atom is accounted for in the chemical structure representations, and whenever possible all byproducts are accounted for, all stoichiometric reagents are correctly balanced, and quantities are entered in a standardised form. Continue reading
Sneak preview: MMDS lookup of assays via OpenPHACTS
 One of the features that’s slated for the next major version of the Mobile Molecular DataSheet (MMDS) is integration with OpenPHACTS pharmacological assay data. The feature is not quite ready yet, but the user interface is taking form, and is almost functional. The basic idea is that you start with a datasheet, with some number of molecules. Then, select the assay lookup feature, and a webservice intermediary cross references each of the molecular structures to the OpenPHACTS service, to pull out every available pharmacology datum, in a form that’s ready to insert into your data sheet as a new column. Looking up the data is quite easy, since OpenPHACTS is preassembled and has a very no-nonsense API, but reconciling a large number of datapoints into one or several columns of numbers is an interesting user interface workflow challenge. Continue reading
One of the features that’s slated for the next major version of the Mobile Molecular DataSheet (MMDS) is integration with OpenPHACTS pharmacological assay data. The feature is not quite ready yet, but the user interface is taking form, and is almost functional. The basic idea is that you start with a datasheet, with some number of molecules. Then, select the assay lookup feature, and a webservice intermediary cross references each of the molecular structures to the OpenPHACTS service, to pull out every available pharmacology datum, in a form that’s ready to insert into your data sheet as a new column. Looking up the data is quite easy, since OpenPHACTS is preassembled and has a very no-nonsense API, but reconciling a large number of datapoints into one or several columns of numbers is an interesting user interface workflow challenge. Continue reading
Lyme disease added to ODDT
 It seemed appropriate to include a new topic to the Open Drug Discovery Teams project: Lyme Disease. After a road trip through New England this summer, seeing signs everywhere warning visitors to be ultra careful not to do anything reckless such as walking anywhere near green foliage, then reading articles about how the disease is actually an underdiagnosed epidemic, it might be time to start thinking of it in the same terms as neglected tropical diseases.
It seemed appropriate to include a new topic to the Open Drug Discovery Teams project: Lyme Disease. After a road trip through New England this summer, seeing signs everywhere warning visitors to be ultra careful not to do anything reckless such as walking anywhere near green foliage, then reading articles about how the disease is actually an underdiagnosed epidemic, it might be time to start thinking of it in the same terms as neglected tropical diseases.
Welcome to the club, ChemDraw Mobile
Yesterday’s buzz in the chemistry software scene revolved around the release of ChemDraw Mobile for iPad (but not iPhone). This is exciting news, even if only for the name: the flagship chemical drawing software from CambridgeSoft has been the go-to brand for creating diagrams for manuscripts for something like a quarter century (give or take). It’s hard to argue that working with chemical structures on mobile devices is an irrelevant hobby when the biggest player in the industry has made the effort to join in, and is giving it top billing with its promotional efforts. Continue reading
Sharing structures with SketchEl and molsync.com
 The latest version of SketchEl (1.60) has added a new feature: sharing of content via molsync.com. From either the molecule editor or the datasheet editor, the underlying content can be directly uploaded, making it openly accessible to the greater internet.
The latest version of SketchEl (1.60) has added a new feature: sharing of content via molsync.com. From either the molecule editor or the datasheet editor, the underlying content can be directly uploaded, making it openly accessible to the greater internet.
While SketchEl is not technically a product of Molecular Materials Informatics, the copyright is held by me, it’s made available via the Gnu Public License, and it gets a bit of maintenance from time to time. It is a conventional and fairly capable 2D structure drawing program that is written in pure Java, and runs on all the major platforms. It can even be used as an applet, though that doesn’t matter as much as it used to. It also allows editing of molecular spreadsheets (“datasheets”) that contain molecules and various scalar data. Over the years (since 2005) it has served as a test bed for a few ideas, like the SketchEl molecule file format, and I still use it regularly for editing structures and collections.
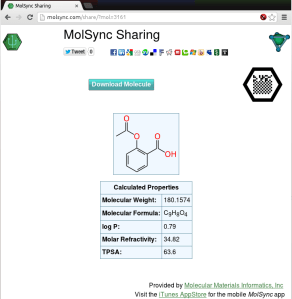
The newly added sharing feature offers to upload the current molecule or datasheet. Once the upload is complete, the result is available in the form of a URL, which can be opened in the browser, e.g. http://molsync.com/share?mol=3161:
This is the same workflow that is available via several mobile apps, like MMDS and MolPrime+, either for sharing directly as a web page, or as a prelude to tweeting out the data. Once you have the data uploaded to the site, the content is served up in viewable form, and can be downloaded in various cheminformatics and graphics formats. It can be shared by any sort of internet distribution means, and there are lots of clickable icons for bouncing it out via Twitter, Facebook, LinkedIn, Google+, etc.
Notice also that the page includes the molecular recognition glyph – that’s the black hexagon with the square dot pattern in the middle. The timing is no coincidence, because it means that as soon as the Living Molecules app gets through the appstore review process, the following workflow will be possible:
- Draw molecules and/or assemble them into a datasheet with textual content, properties, etc., using any desktop platform (Linux, Windows, Mac, whatever)
- Share the content on molsync.com
- Export the molecular glyph and include the graphic on a poster
- Put up the poster at a conference, and anyone with an iThing can snap the glyph and access the underlying data
Of course, using many of the content creation apps the data can be created on a mobile device as well, but adding the feature to SketchEl now makes that a choice: create content on mobile or desktop, whichever is most convenient at the time. And it’s not just for making posters either, but that is one of the original use case proposals.
Molecular glyphs on MolSync.com
 The molsync.com sharing page has a new feature, as shown on the screenshot to the right (click on it to see the live version): there’s now a black hexagon with a grid pattern in the middle, which is a molecular recognition glyph designed to work with the Living Molecules app. The simplest way to describe it is that it’s a molecular QR code, suitable for inclusion on posters or various other documents. The content can be accessed by using the Living Molecules app to “snap” the glyph, after which it continues on to download the data and make it available for viewing, storing and exporting.
The molsync.com sharing page has a new feature, as shown on the screenshot to the right (click on it to see the live version): there’s now a black hexagon with a grid pattern in the middle, which is a molecular recognition glyph designed to work with the Living Molecules app. The simplest way to describe it is that it’s a molecular QR code, suitable for inclusion on posters or various other documents. The content can be accessed by using the Living Molecules app to “snap” the glyph, after which it continues on to download the data and make it available for viewing, storing and exporting.
The app has been submitted to the iTunes AppStore, so with any luck, it will be available for download (free) in a week or so!