 In the previous post, I introduced the BioAssay Express project, which is based on the idea that when assay protocols are described using semantic annotations rather than just scientific text, a whole realm of software analysis becomes possible. Whenever there’s a bioassay experiment protocol at hand, chances are there are also some molecules with activity measurements not far away: which is the subject of this article.
In the previous post, I introduced the BioAssay Express project, which is based on the idea that when assay protocols are described using semantic annotations rather than just scientific text, a whole realm of software analysis becomes possible. Whenever there’s a bioassay experiment protocol at hand, chances are there are also some molecules with activity measurements not far away: which is the subject of this article.
The raw data from which the BioAssay Express draws is currently entirely from PubChem, on account of a large amount of data being stored in one place and accessible via one API. Unlike other open repositories (such as ChEMBL), many of the assays have the full text description of the experimental protocol included in the record, which essential for our purposes; and fortunately, PubChem keeps track of who uploaded each assay, which happens to have a near perfect correlation with whether a given assay is suitable.
PubChem is of course more famous for being a giant database of molecules, and many of these are connected to one or more assays via measurement data. Since it went online, the BioAssay Express project has been regularly interrogating the PubChem assay collection, looking for new entries provided by one of the whitelisted contributors. Since last week, this background task has been augmented by two more, which keep themselves busy downloading the measurements for each assay, and for each of these, making sure that all the compounds have structures.
The first visible consequence of this internal expansion is another section at the bottom of the assay assignment page, which you can find on pages such as: http://www.bioassayexpress.com/BioAssayExpress/assign.jsp?pubchemAID=1599:
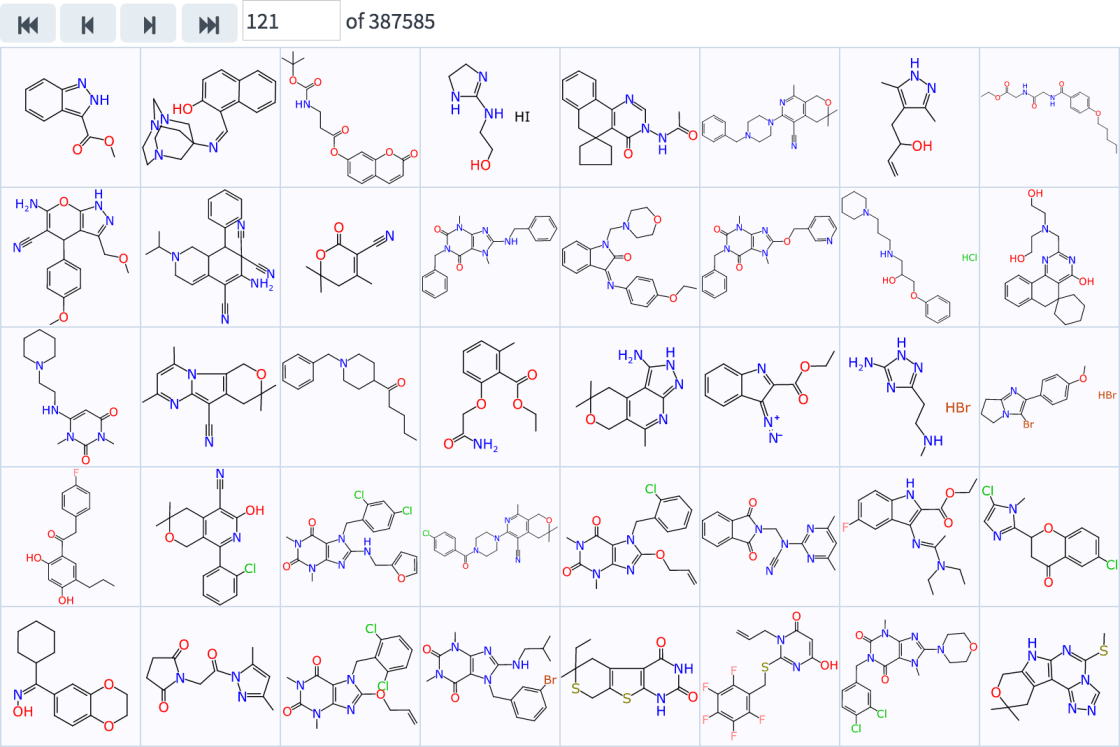
Right now the grid view is not hugely much more functional than it looks: you can step through one page at a time, or click on a compound for more information, which is currently expressed as a detail dialog:
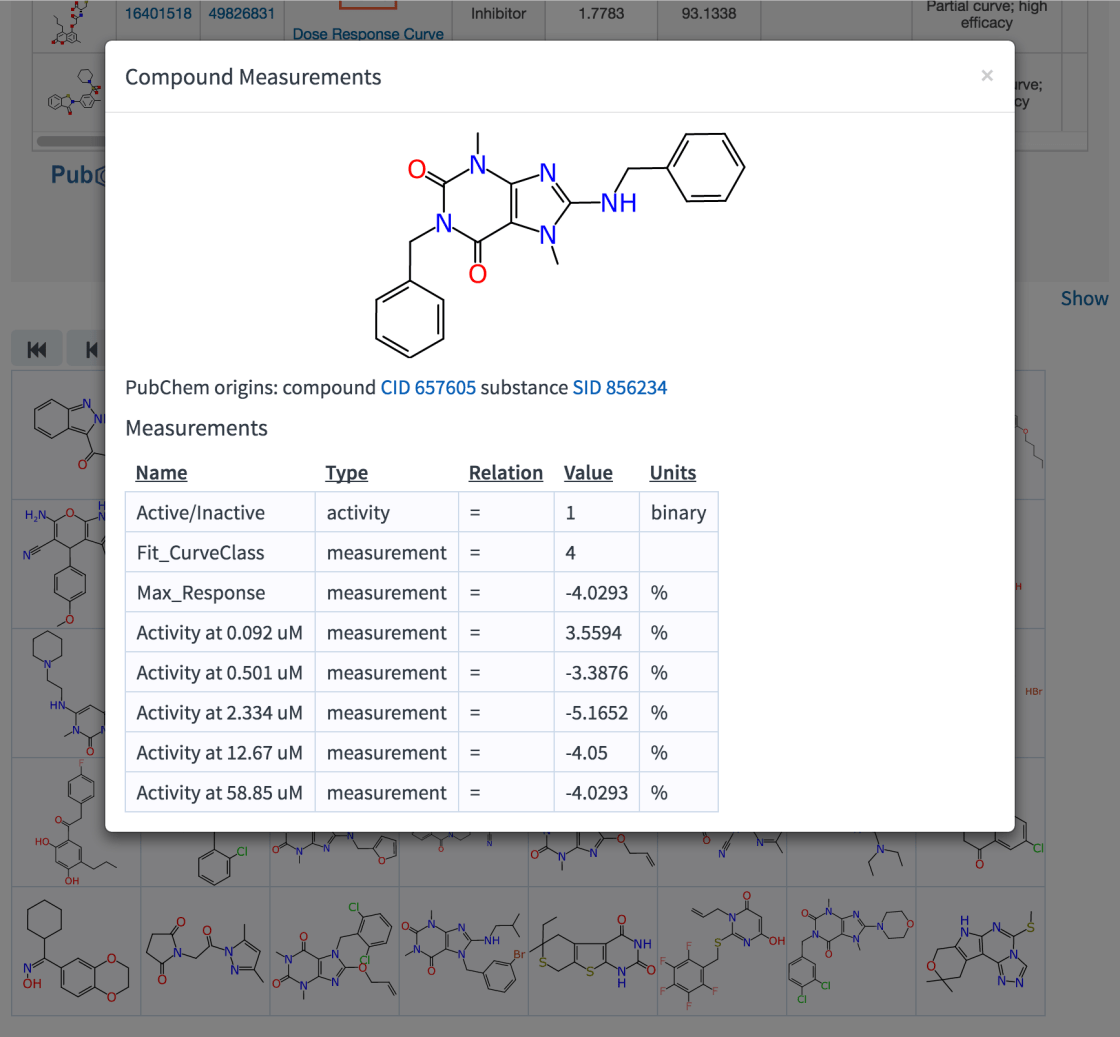
What really is exciting about this – and will become more apparent in the near future – is how these structures made the journey from a Molfile record stored on the server to your browser. If you were to guess that these were bitmapped PNG files generated by server and handed off down the wire to be rendered as-is, you would be mistaken. They are actually vector graphics images, displayed using the SVG format (meaning that they look perfectly beautiful at any resolution). That in itself would be oh so very 2013, except they’re not just any embedded SVG drawings, they are generated by the browser using a toolkit that is running in JavaScript without any help from a server.
This is possible because the toolkit that I’ve been working on under the banner of Molecular Materials Informatics for some years has hopped across a number of platforms. The main reference implementation is written in regular Java, and requisite pieces have been migrated to Android/BlackBerry Java, Objective-C, Swift, and most recently TypeScript/JavaScript. The activity several months ago with the MolSync(.com) site involved a hybrid mix of server & client cheminformatics, but the toolkit has been pushed further: a bunch of important functionality can be run without any server dependence, and amongst these are the algorithms needed to turn a 2D sketch (encoded in a cheminformatics format, like MDL Molfile) into a fully rendered image that is truly publication quality. This is one of those tasks that is easy to do badly and hard to do well, like so much of cheminformatics: I published an article about it in 2013 (entitled Rendering Molecular Sketches for Publication Quality Output if you want to know just how much work is involved). Now the entire algorithm is coded up in TypeScript/JavaScript, and it can be plugged into any web application. The functionality is being augmented rapidly, and the next frontier is to completely port the sketcher: most of the algorithms have been translated to the client side, i.e. they are now happening on the web without a server, but there are few more leftovers to migrate before sketching is completely server-independent.
In addition to powering the molsync.com site, and now being added to the functionality of the BioAssay Express, this toolkit (“WebMolKit”) is actually open source. Right now the source code is on a private GitHub account, which will be opened up to the public once it has had a little bit more time to mature (it’s very green at this moment). The license permits anyone to use it under the Gnu Public License, as soon as the switch is flipped and the code is made available.
From the point of view of the BioAssay Express, adding molecules to applicable pages is a nice novelty feature, though it is not so amazing unto itself since we already have the use of PubChem‘s embeddable widget which achieves much the same effect. The real value kicks in once it is possible to actually do things with the molecules, and the next step along that road is to allow some reorganising of the measurement columns (e.g. pick the activity measurement you want and rename it to “IC50”), and then be able to hit a download button and grab the entire thing as a single SDfile, with all the molecules, and just the columns you asked for. Right now that particular step is quite difficult to accomplish using raw PubChem access, and takes a fair bit of cheminformatics know-how.
But even that is a relatively minor step in the greater scheme of things: it gets really interesting when this functionality is connected up to features that return more than one assay (e.g. the Browse Assays feature). Right now this is a great way to find assays according to certain criteria, but in the not too distant future, it will also be able to show a union of all the molecules for all the selected assays. They will generally have a heterogeneous collection of measurements, which will be able to be selected, renamed and filtered, prior to downloading the whole thing as one big SAR dataset. This workflow is a pretty useful way to go from a vague idea to a dataset that is ready to be plugged into a model building exercise, or a visualisation tool.
If this sounds interesting, then keep an eye on the BioAssay Express site, the Collaborative Drug Discovery home page, and announcements on this blog.