Using chemical substructures with query annotations is a cheminformatics trick that goes back a long way: by annotating the query structure with additional logical constraints, the utility of this functionality can be juiced up, by matching a variety of certain kinds of fragments, rather than literal pieces. The amount of work involved to implement such a system is quite evenly divided between providing the low-level algorithm for doing the matching, and providing a high-level editor for people to design the queries. One of the “TO-DO” features in the XMDS beta version has taken some steps toward fulfilling the latter.
Using chemical substructures with query annotations is a cheminformatics trick that goes back a long way: by annotating the query structure with additional logical constraints, the utility of this functionality can be juiced up, by matching a variety of certain kinds of fragments, rather than literal pieces. The amount of work involved to implement such a system is quite evenly divided between providing the low-level algorithm for doing the matching, and providing a high-level editor for people to design the queries. One of the “TO-DO” features in the XMDS beta version has taken some steps toward fulfilling the latter.
Last year, one of the interesting projects I worked on is the Medicinal Chemistry Toolkit app on behalf of the Royal Society of Chemistry, which had a challenging requirement: the Astra-Zeneca structure filters needed to be provided as a realtime annotation, and the app was not allowed to call out to a webservice. This basically meant that a lot of substructure queries needed to be implemented on mobile. At the time, my company’s com.mmi back-end software stack (written in Java) had a competent substructure search implementation, but it didn’t support all of the query features that are available within a complete implementation of SMARTS queries, which can get really seriously gnarly when you bring in sub-fragment query matches of unlimited depth. Which the Astra-Zeneca filters include a lot of.
Upgrading the SketchEl format to include query features was all in a day’s work, and adding these features into the Java-based cheminformatics toolkit was not unreasonably challenging. The subfragments took a bit of finesse, but ultimately the implementation turned out to be quite clean – it is recursive afterall, and with a few little tweaks to keep performance within reason, this challenge folded fairly quickly.
The harder part was actually rebuilding the queries, using SMARTS and human “readable” description as the basis, but not the definitive answer, and expressing these in the sketch-like query format that is query-extended SketchEl. These got a little too complex to do manually, and I ended up resorting to something that I prefer not to do: making a somewhat hacky extension to an existing software package to help with a singular task. This started by adding query property editor dialogs for atoms and bonds to the SketchEl open source editor:
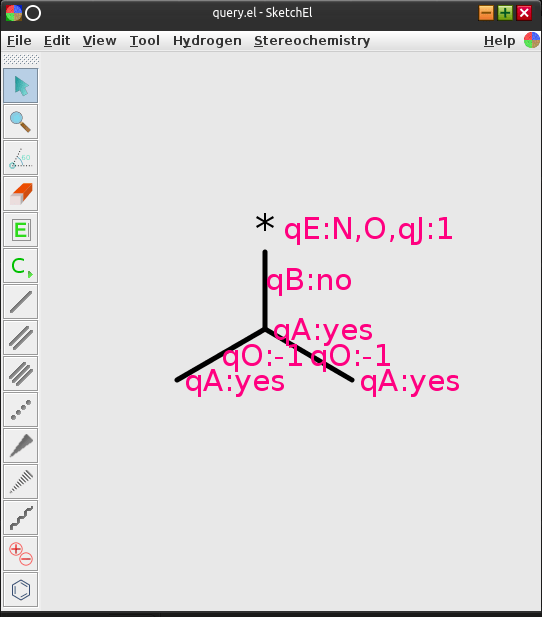
These dialogs are in no danger of winning any design awards, but the actual display of the annotated properties is even uglier:
Aesthetics aside though, it works: the “*” atom is a terminal N or O, emerging from an aromatic ring system, in this example. The query is overspecified, because this is a good thing (it gives the query matcher more ways to fast-fail). The underlying SketchEl molecule representation is pleasantly simple (for cheminformatics nerds at least):
SketchEl!(4,3) *=-2.0500,5.3500;0,0,i0,qE:N\002CO,qJ:1 C=-2.0500,3.8500;0,0,i1,qA:yes C=-3.3490,3.1000;0,0,i3,qA:yes C=-0.7510,3.1000;0,0,i3,qA:yes 1-2=1,0,qB:no 2-3=1,0,qO:-1 2-4=1,0,qO:-1 !End
The augmentation of the SketchEl program also included a way to subsume fragments to use as recursive sub-fragment matches: a menu item to include or exclude previously drawn components, and wrap them in as inline structures. This is roughly equivalent to people writing cryptic SMARTS queries like [C,N;$(C=N(C)c),!$(C=O)] (imagine something like that but 500 characters long). The kludgey extension to SketchEl is a delightful user experience by comparison, as hoary as it is.
That was good enough to get the Astra-Zeneca filters working adequately, and to port the algorithm to mobile. But on the to-do list has been the creation of a somewhat nicer editing feature, for the OS X Molecular DataSheet (XMDS) app for Mac. The atom and bond query features are now implemented within the atom- and bond-editing panels:
In this implementation, the pertinent features are added one at a time, rather than one big dialog with all options shown. The actual display of query features on the molecule is still rather ugly: this is a placeholder. While SketchEl will not be getting a beautification update anytime soon, this is on the menu for XMDS, but in the interests of getting it operational, this part of the visuals will have to wait.
But consider the atom query feature editor when subfragments are involved:
The XMDS app pulls out the fragment (include/exclude) pieces and renders them, and also indicates the atom that overlaps with the current query atom. The rendition is very similar to the way the abbreviation editor works.
There is a real reason why I’m back to working on substructure query features, but that will have to wait for another post.